
Single-stranded binding protein
| SSB | |||||||||
|---|---|---|---|---|---|---|---|---|---|
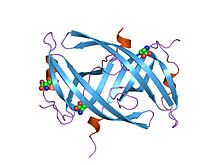 Crystal structure of PriB- a primosomal DNA replication protein of Escherichia coli
| |||||||||
| Identifiers | |||||||||
| Symbol | SSB | ||||||||
| Pfam | PF00436 | ||||||||
| Pfam clan | CL0021 | ||||||||
| InterPro | IPR000424 | ||||||||
| PROSITE | PDOC00602 | ||||||||
| SCOP2 | 1kaw / SCOPe / SUPFAM | ||||||||
| TCDB | 3.A.7 | ||||||||
| |||||||||
Single-stranded binding proteins (SSBs) are a class of proteins that have been identified in both viruses and organisms from bacteria to humans.
Viral SSB
| Viral_DNA_bp | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Single stranded DNA-binding protein(icp8) from herpes simplex virus-1
| |||||||||
| Identifiers | |||||||||
| Symbol | Viral_DNA_bp | ||||||||
| Pfam | PF00747 | ||||||||
| InterPro | IPR000635 | ||||||||
| |||||||||
Although the overall picture of human cytomegalovirus (HHV-5) DNA synthesis appears typical of the herpesviruses, some novel features are emerging.
Structure
In ICP8, the herpes simplex virus (HSV-1) single-strand DNA-binding protein (ssDNA-binding protein (SSB)), the head consists of the eight alpha helices. The front side of the neck region consists of a five-stranded beta-sheet and two alpha helices, whereas the back side is a three-stranded beta-sheet The shoulder part of the N-terminal domain contains an alpha-helical and beta-sheet region. The herpes simplex virus (HSV-1) SSB, ICP8, is a nuclear protein that, along other replication proteins is required for viral DNA replication during lytic infection.
Mechanism
Six herpes virus-group-common genes encode proteins that likely constitute the replication fork machinery, including a two-subunit DNA polymerase, a Helicase-primase complex and a single-stranded DNA-binding protein. The human herpesvirus 1 (HHV-1) single-strand DNA-binding protein ICP8 is a 128kDa zinc metalloprotein. Photoaffinity labeling has shown that the region encompassing amino acid residues 368-902 contains the single-strand DNA-binding site of ICP8. The HHHV-1 UL5, UL8, and UL52 genes encode an essential heterotrimeric DNA helicase-primase that is responsible for concomitant DNA unwinding and primer synthesis at the viral DNA replication fork. ICP8 may stimulate DNA unwinding and enable bypass of cisplatin damaged DNA by recruiting the helicase-primase to the DNA.
Bacterial SSB
SSB protein domains in bacteria are important maintaining DNA metabolism, more specifically DNA replication, repair and recombination. It has a structure of three beta-strands to a single six-stranded beta-sheet to form a dimer.
Eukaryotic replication protein A
| Replication protein A | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| (heterotrimer) | |||||||||||||
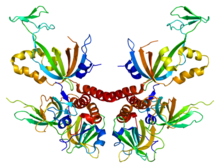 This is an image of human Replication protein A. From PDB: 1L1O Proteopedia protein A Replication protein A
| |||||||||||||
| Function | damaged DNA binding, single-stranded DNA binding | ||||||||||||
| |||||||||||||
Replication protein A is the functional equivalent of SSB in the nucleus of eukaryotic cells, though there is no sequence homology.
Eukaryotic mitochondrial SSB
The mitochondria of eukaryotic cells contain their own single stranded DNA binding protein. Human mitochondrial SSB (mtSSB) binds to single-stranded mitochondrial DNA as a tetramer and has sequence similarity to bacterial SSB. Human mtSSB is encoded by the SSBP1 gene. In yeast, it is encoded by the RIM1 gene.
Role in Genome Repair and Anti-aging
Recently, it has been found that it 1. Helps protect the genome, 2. Is vital for stem cells and 3. Is involved with maintaining telomere length.
See also
External links
- Single-Stranded+DNA+Binding+Proteins at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- SSB in PFAM
| Initiation |
|
||||||
|---|---|---|---|---|---|---|---|
| Replication |
|
||||||
| Termination | |||||||