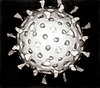
Bacteriophage

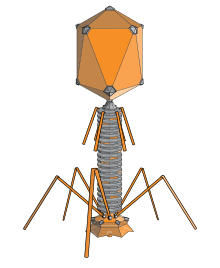

A bacteriophage (/bækˈtɪərioʊfeɪdʒ/), also known informally as a phage (/ˈfeɪdʒ/), is a duplodnaviria virus that infects and replicates within bacteria and archaea. The term was derived from "bacteria" and the Greek φαγεῖν (phagein), meaning "to devour". Bacteriophages are composed of proteins that encapsulate a DNA or RNA genome, and may have structures that are either simple or elaborate. Their genomes may encode as few as four genes (e.g. MS2) and as many as hundreds of genes. Phages replicate within the bacterium following the injection of their genome into its cytoplasm.
Bacteriophages are among the most common and diverse entities in the biosphere. Bacteriophages are ubiquitous viruses, found wherever bacteria exist. It is estimated there are more than 1031 bacteriophages on the planet, more than every other organism on Earth, including bacteria, combined. Viruses are the most abundant biological entity in the water column of the world's oceans, and the second largest component of biomass after prokaryotes, where up to 9x108virions per millilitre have been found in microbial mats at the surface, and up to 70% of marine bacteria may be infected by phages.
Phages have been used since the late 20th century as an alternative to antibiotics in the former Soviet Union and Central Europe, as well as in France. They are seen as a possible therapy against multi-drug-resistant strains of many bacteria (see phage therapy).
Phages are known to interact with the immune system both indirectly via bacterial expression of phage-encoded proteins and directly by influencing innate immunity and bacterial clearance. Phage–host interactions are becoming increasingly important areas of research.
Classification
Bacteriophages occur abundantly in the biosphere, with different genomes and lifestyles. Phages are classified by the International Committee on Taxonomy of Viruses (ICTV) according to morphology and nucleic acid.


| Order | Family | Morphology | Nucleic acid | Examples |
|---|---|---|---|---|
| Belfryvirales | Turriviridae | Enveloped, isometric | Linear dsDNA | |
| Caudovirales | Ackermannviridae | Nonenveloped, contractile tail | Linear dsDNA | |
| Autographiviridae | Nonenveloped, noncontractile tail (short) | Linear dsDNA | ||
| Chaseviridae | Linear dsDNA | |||
| Demerecviridae | Linear dsDNA | |||
| Drexlerviridae | Linear dsDNA | |||
| Guenliviridae | Linear dsDNA | |||
| Herelleviridae | Nonenveloped, contractile tail | Linear dsDNA | ||
| Myoviridae | Nonenveloped, contractile tail | Linear dsDNA | T4, Mu, P1, P2 | |
| Siphoviridae | Nonenveloped, noncontractile tail (long) | Linear dsDNA | λ, T5, HK97, N15 | |
| Podoviridae | Nonenveloped, noncontractile tail (short) | Linear dsDNA | T7, T3, Φ29, P22 | |
| Rountreeviridae | Linear dsDNA | |||
| Salasmaviridae | Linear dsDNA | |||
| Schitoviridae | Linear dsDNA | |||
| Zobellviridae | Linear dsDNA | |||
| Halopanivirales | Sphaerolipoviridae | Enveloped, isometric | Linear dsDNA | |
| Simuloviridae | Enveloped, isometric | Linear dsDNA | ||
| Matshushitaviridae | Enveloped, isometric | Linear dsDNA | ||
| Haloruvirales | Pleolipoviridae | Enveloped, pleomorphic | Circular ssDNA, circular dsDNA, or linear dsDNA | |
| Kalamavirales | Tectiviridae | Nonenveloped, isometric | Linear dsDNA | |
| Ligamenvirales | Lipothrixviridae | Enveloped, rod-shaped | Linear dsDNA | Acidianus filamentous virus 1 |
| Rudiviridae | Nonenveloped, rod-shaped | Linear dsDNA | Sulfolobus islandicus rod-shaped virus 1 | |
| Mindivirales | Cystoviridae | Enveloped, spherical | Linear dsRNA | Φ6 |
| Norzivirales | Atkinsviridae | Nonenveloped, isometric | Linear ssRNA | |
| Duinviridae | Nonenveloped, isometric | Linear ssRNA | ||
| Fiersviridae | Nonenveloped, isometric | Linear ssRNA | MS2, Qβ | |
| Solspiviridae | Nonenveloped, isometric | Linear ssRNA | ||
| Petitvirales | Microviridae | Nonenveloped, isometric | Circular ssDNA | ΦX174 |
| Primavirales | Tristromaviridae | Enveloped, rod-shaped | Linear dsDNA | |
| Timlovirales | Blumeviridae | Nonenveloped, isometric | Linear ssRNA | |
| Steitzviridae | Nonenveloped, isometric | Linear ssRNA | ||
| Tubulavirales | Inoviridae | Nonenveloped, filamentous | Circular ssDNA | M13 |
| Paulinoviridae | Nonenveloped, filamentous | Circular ssDNA | ||
| Plectroviridae | Nonenveloped, filamentous | Circular ssDNA | ||
| Vinavirales | Corticoviridae | Nonenveloped, isometric | Circular dsDNA | PM2 |
| Durnavirales | Picobirnaviridae (proposal) | Nonenveloped, isometric | Linear dsRNA | |
| Unassigned | Ampullaviridae | Enveloped, bottle-shaped | Linear dsDNA | |
| Autolykiviridae | Nonenveloped, isometric | Linear dsDNA | ||
| Bicaudaviridae | Nonenveloped, lemon-shaped | Circular dsDNA | ||
| Clavaviridae | Nonenveloped, rod-shaped | Circular dsDNA | ||
| Finnlakeviridae | Nonenveloped, isometric | Circular ssDNA | FLiP | |
| Fuselloviridae | Nonenveloped, lemon-shaped | Circular dsDNA | ||
| Globuloviridae | Enveloped, isometric | Linear dsDNA | ||
| Guttaviridae | Nonenveloped, ovoid | Circular dsDNA | ||
| Halspiviridae | Nonenveloped, lemon-shaped | Linear dsDNA | ||
| Plasmaviridae | Enveloped, pleomorphic | Circular dsDNA | ||
| Portogloboviridae | Enveloped, isometric | Circular dsDNA | ||
| Thaspiviridae | Nonenveloped, lemon-shaped | Linear dsDNA | ||
| Spiraviridae | Nonnveloped, rod-shaped | Circular ssDNA |
It has been suggested that members of Picobirnaviridae infect bacteria, but not mammals.
There are also many unassigned genera of the class Leviviricetes: Chimpavirus, Hohglivirus, Mahrahvirus, Meihzavirus, Nicedsevirus, Sculuvirus, Skrubnovirus, Tetipavirus and Winunavirus containing linear ssRNA genomes and the unassigned genus Lilyvirus of the order Caudovirales containing a linear dsDNA genome.
History
In 1896, Ernest Hanbury Hankin reported that something in the waters of the Ganges and Yamuna rivers in India had a marked antibacterial action against cholera and it could pass through a very fine porcelain filter. In 1915, British bacteriologist Frederick Twort, superintendent of the Brown Institution of London, discovered a small agent that infected and killed bacteria. He believed the agent must be one of the following:
- a stage in the life cycle of the bacteria
- an enzyme produced by the bacteria themselves, or
- a virus that grew on and destroyed the bacteria
Twort's research was interrupted by the onset of World War I, as well as a shortage of funding and the discoveries of antibiotics.
Independently, French-Canadian microbiologist Félix d'Hérelle, working at the Pasteur Institute in Paris, announced on 3 September 1917 that he had discovered "an invisible, antagonistic microbe of the dysentery bacillus". For d'Hérelle, there was no question as to the nature of his discovery: "In a flash I had understood: what caused my clear spots was in fact an invisible microbe... a virus parasitic on bacteria." D'Hérelle called the virus a bacteriophage, a bacteria-eater (from the Greek phagein, meaning "to devour"). He also recorded a dramatic account of a man suffering from dysentery who was restored to good health by the bacteriophages. It was d'Hérelle who conducted much research into bacteriophages and introduced the concept of phage therapy.
Nobel prizes awarded for phage research
In 1969, Max Delbrück, Alfred Hershey, and Salvador Luria were awarded the Nobel Prize in Physiology or Medicine for their discoveries of the replication of viruses and their genetic structure. Specifically the work of Hershey, as contributor to the Hershey–Chase experiment in 1952 provided convincing evidence that DNA, not protein, was the genetic material of life. Delbrück and Luria carried out the Luria–Delbrück experiment which demonstrated statistically that mutations in bacteria occur randomly and thus follow Darwinian rather than Lamarckian principles.
Uses
Phage therapy
Phages were discovered to be antibacterial agents and were used in the former Soviet Republic of Georgia (pioneered there by Giorgi Eliava with help from the co-discoverer of bacteriophages, Félix d'Hérelle) during the 1920s and 1930s for treating bacterial infections. They had widespread use, including treatment of soldiers in the Red Army. However, they were abandoned for general use in the West for several reasons:
- Antibiotics were discovered and marketed widely. They were easier to make, store, and prescribe.
- Medical trials of phages were carried out, but a basic lack of understanding of phages raised questions about the validity of these trials.
- Publication of research in the Soviet Union was mainly in the Russian or Georgian languages and for many years, was not followed internationally.
The use of phages has continued since the end of the Cold War in Russia, Georgia, and elsewhere in Central and Eastern Europe. The first regulated, randomized, double-blind clinical trial was reported in the Journal of Wound Care in June 2009, which evaluated the safety and efficacy of a bacteriophage cocktail to treat infected venous ulcers of the leg in human patients. The FDA approved the study as a Phase I clinical trial. The study's results demonstrated the safety of therapeutic application of bacteriophages, but did not show efficacy. The authors explained that the use of certain chemicals that are part of standard wound care (e.g. lactoferrin or silver) may have interfered with bacteriophage viability. Shortly after that, another controlled clinical trial in Western Europe (treatment of ear infections caused by Pseudomonas aeruginosa) was reported in the journal Clinical Otolaryngology in August 2009. The study concludes that bacteriophage preparations were safe and effective for treatment of chronic ear infections in humans. Additionally, there have been numerous animal and other experimental clinical trials evaluating the efficacy of bacteriophages for various diseases, such as infected burns and wounds, and cystic fibrosis-associated lung infections, among others. On the other hand, phages of Inoviridae have been shown to complicate biofilms involved in pneumonia and cystic fibrosis and to shelter the bacteria from drugs meant to eradicate disease, thus promoting persistent infection.
Meanwhile, bacteriophage researchers have been developing engineered viruses to overcome antibiotic resistance, and engineering the phage genes responsible for coding enzymes that degrade the biofilm matrix, phage structural proteins, and the enzymes responsible for lysis of the bacterial cell wall. There have been results showing that T4 phages that are small in size and short-tailed can be helpful in detecting E. coli in the human body.
Therapeutic efficacy of a phage cocktail was evaluated in a mice model with nasal infection of multidrug-resistant (MDR) A. baumannii. Mice treated with the phage cocktail showed a 2.3-fold higher survival rate compared to those untreated at seven days post-infection. In 2017, a patient with a pancreas compromised by MDR A. baumannii was put on several antibiotics; despite this, the patient's health continued to deteriorate during a four-month period. Without effective antibiotics, the patient was subjected to phage therapy using a phage cocktail containing nine different phages that had been demonstrated to be effective against MDR A. baumannii. Once on this therapy the patient's downward clinical trajectory reversed, and returned to health.
D'Herelle "quickly learned that bacteriophages are found wherever bacteria thrive: in sewers, in rivers that catch waste runoff from pipes, and in the stools of convalescent patients." This includes rivers traditionally thought to have healing powers, including India's Ganges River.
Other
Food industry – Phages have increasingly been used to safen food products and to forestall spoilage bacteria. Since 2006, the United States Food and Drug Administration (FDA) and United States Department of Agriculture (USDA) have approved several bacteriophage products. LMP-102 (Intralytix) was approved for treating ready-to-eat (RTE) poultry and meat products. In that same year, the FDA approved LISTEX (developed and produced by Micreos) using bacteriophages on cheese to kill Listeria monocytogenes bacteria, in order to give them generally recognized as safe (GRAS) status. In July 2007, the same bacteriophage were approved for use on all food products. In 2011 USDA confirmed that LISTEX is a clean label processing aid and is included in USDA. Research in the field of food safety is continuing to see if lytic phages are a viable option to control other food-borne pathogens in various food products.
Diagnostics – In 2011, the FDA cleared the first bacteriophage-based product for in vitro diagnostic use. The KeyPath MRSA/MSSA Blood Culture Test uses a cocktail of bacteriophage to detect Staphylococcus aureus in positive blood cultures and determine methicillin resistance or susceptibility. The test returns results in about five hours, compared to two to three days for standard microbial identification and susceptibility test methods. It was the first accelerated antibiotic-susceptibility test approved by the FDA.
Counteracting bioweapons and toxins – Government agencies in the West have for several years been looking to Georgia and the former Soviet Union for help with exploiting phages for counteracting bioweapons and toxins, such as anthrax and botulism. Developments are continuing among research groups in the U.S. Other uses include spray application in horticulture for protecting plants and vegetable produce from decay and the spread of bacterial disease. Other applications for bacteriophages are as biocides for environmental surfaces, e.g., in hospitals, and as preventative treatments for catheters and medical devices before use in clinical settings. The technology for phages to be applied to dry surfaces, e.g., uniforms, curtains, or even sutures for surgery now exists. Clinical trials reported in Clinical Otolaryngology show success in veterinary treatment of pet dogs with otitis.
The SEPTIC bacterium sensing and identification method uses the ion emission and its dynamics during phage infection and offers high specificity and speed for detection.
Phage display is a different use of phages involving a library of phages with a variable peptide linked to a surface protein. Each phage genome encodes the variant of the protein displayed on its surface (hence the name), providing a link between the peptide variant and its encoding gene. Variant phages from the library may be selected through their binding affinity to an immobilized molecule (e.g., botulism toxin) to neutralize it. The bound, selected phages can be multiplied by reinfecting a susceptible bacterial strain, thus allowing them to retrieve the peptides encoded in them for further study.
Antimicrobial drug discovery – Phage proteins often have antimicrobial activity and may serve as leads for peptidomimetics, i.e. drugs that mimic peptides.Phage-ligand technology makes use of phage proteins for various applications, such as binding of bacteria and bacterial components (e.g. endotoxin) and lysis of bacteria.
Basic research – Bacteriophages are important model organisms for studying principles of evolution and ecology.
Detriments
Dairy industry
Bacteriophages present in the environment can cause cheese to not ferment. In order to avoid this, mixed-strain starter cultures and culture rotation regimes can be used.Genetic engineering of culture microbes – especially Lactococcus lactis and Streptococcus thermophilus – have been studied for genetic analysis and modification to improve phage resistance. This has especially focused on plasmid and recombinant chromosomal modifications.
Some research has focused on the potential of bacteriophages as antimicrobial against foodborne pathogens and biofilm formation within the dairy industry. As the spread of antibiotic resistance is a main concern within the dairy industry, phages can serve as a promising alternative.
Replication
The life cycle of bacteriophages tends to be either a lytic cycle or a lysogenic cycle. In addition, some phages display pseudolysogenic behaviors.
With lytic phages such as the T4 phage, bacterial cells are broken open (lysed) and destroyed after immediate replication of the virion. As soon as the cell is destroyed, the phage progeny can find new hosts to infect. Lytic phages are more suitable for phage therapy. Some lytic phages undergo a phenomenon known as lysis inhibition, where completed phage progeny will not immediately lyse out of the cell if extracellular phage concentrations are high. This mechanism is not identical to that of the temperate phage going dormant and usually is temporary.
In contrast, the lysogenic cycle does not result in immediate lysing of the host cell. Those phages able to undergo lysogeny are known as temperate phages. Their viral genome will integrate with host DNA and replicate along with it, relatively harmlessly, or may even become established as a plasmid. The virus remains dormant until host conditions deteriorate, perhaps due to depletion of nutrients, then, the endogenous phages (known as prophages) become active. At this point they initiate the reproductive cycle, resulting in lysis of the host cell. As the lysogenic cycle allows the host cell to continue to survive and reproduce, the virus is replicated in all offspring of the cell. An example of a bacteriophage known to follow the lysogenic cycle and the lytic cycle is the phage lambda of E. coli.
Sometimes prophages may provide benefits to the host bacterium while they are dormant by adding new functions to the bacterial genome, in a phenomenon called lysogenic conversion. Examples are the conversion of harmless strains of Corynebacterium diphtheriae or Vibrio cholerae by bacteriophages, to highly virulent ones that cause diphtheria or cholera, respectively. Strategies to combat certain bacterial infections by targeting these toxin-encoding prophages have been proposed.
Attachment and penetration
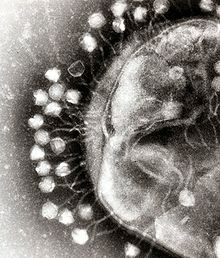
Bacterial cells are protected by a cell wall of polysaccharides, which are important virulence factors protecting bacterial cells against both immune host defenses and antibiotics.To enter a host cell, bacteriophages bind to specific receptors on the surface of bacteria, including lipopolysaccharides, teichoic acids, proteins, or even flagella. This specificity means a bacteriophage can infect only certain bacteria bearing receptors to which they can bind, which in turn, determines the phage's host range. Polysaccharide-degrading enzymes are virion-associated proteins that enzymatically degrade the capsular outer layer of their hosts at the initial step of a tightly programmed phage infection process. Host growth conditions also influence the ability of the phage to attach and invade them. As phage virions do not move independently, they must rely on random encounters with the correct receptors when in solution, such as blood, lymphatic circulation, irrigation, soil water, etc.
Myovirus bacteriophages use a hypodermic syringe-like motion to inject their genetic material into the cell. After contacting the appropriate receptor, the tail fibers flex to bring the base plate closer to the surface of the cell. This is known as reversible binding. Once attached completely, irreversible binding is initiated and the tail contracts, possibly with the help of ATP present in the tail, injecting genetic material through the bacterial membrane. The injection is accomplished through a sort of bending motion in the shaft by going to the side, contracting closer to the cell and pushing back up. Podoviruses lack an elongated tail sheath like that of a myovirus, so instead, they use their small, tooth-like tail fibers enzymatically to degrade a portion of the cell membrane before inserting their genetic material.
Synthesis of proteins and nucleic acid
Within minutes, bacterial ribosomes start translating viral mRNA into protein. For RNA-based phages, RNA replicase is synthesized early in the process. Proteins modify the bacterial RNA polymerase so it preferentially transcribes viral mRNA. The host's normal synthesis of proteins and nucleic acids is disrupted, and it is forced to manufacture viral products instead. These products go on to become part of new virions within the cell, helper proteins that contribute to the assemblage of new virions, or proteins involved in cell lysis. In 1972, Walter Fiers (University of Ghent, Belgium) was the first to establish the complete nucleotide sequence of a gene and in 1976, of the viral genome of bacteriophage MS2. Some dsDNA bacteriophages encode ribosomal proteins, which are thought to modulate protein translation during phage infection.
Virion assembly
In the case of the T4 phage, the construction of new virus particles involves the assistance of helper proteins that act catalytically during phage morphogenesis. The base plates are assembled first, with the tails being built upon them afterward. The head capsids, constructed separately, will spontaneously assemble with the tails. During assembly of the phage T4 virion, the morphogenetic proteins encoded by the phage genes interact with each other in a characteristic sequence. Maintaining an appropriate balance in the amounts of each of these proteins produced during viral infection appears to be critical for normal phage T4 morphogenesis. The DNA is packed efficiently within the heads. The whole process takes about 15 minutes.
Release of virions
Phages may be released via cell lysis, by extrusion, or, in a few cases, by budding. Lysis, by tailed phages, is achieved by an enzyme called endolysin, which attacks and breaks down the cell wall peptidoglycan. An altogether different phage type, the filamentous phage, make the host cell continually secrete new virus particles. Released virions are described as free, and, unless defective, are capable of infecting a new bacterium. Budding is associated with certain Mycoplasma phages. In contrast to virion release, phages displaying a lysogenic cycle do not kill the host and instead become long-term residents as prophages.
Communication
Research in 2017 revealed that the bacteriophage Φ3T makes a short viral protein that signals other bacteriophages to lie dormant instead of killing the host bacterium. Arbitrium is the name given to this protein by the researchers who discovered it.
Genome structure
Given the millions of different phages in the environment, phage genomes come in a variety of forms and sizes. RNA phages such as MS2 have the smallest genomes, with only a few kilobases. However, some DNA phages such as T4 may have large genomes with hundreds of genes; the size and shape of the capsid varies along with the size of the genome. The largest bacteriophage genomes reach a size of 735 kb.

Bacteriophage genomes can be highly mosaic, i.e. the genome of many phage species appear to be composed of numerous individual modules. These modules may be found in other phage species in different arrangements. Mycobacteriophages, bacteriophages with mycobacterial hosts, have provided excellent examples of this mosaicism. In these mycobacteriophages, genetic assortment may be the result of repeated instances of site-specific recombination and illegitimate recombination (the result of phage genome acquisition of bacterial host genetic sequences). Evolutionary mechanisms shaping the genomes of bacterial viruses vary between different families and depend upon the type of the nucleic acid, characteristics of the virion structure, as well as the mode of the viral life cycle.
Some marine roseobacter phages contain deoxyuridine (dU) instead of deoxythymidine (dT) in their genomic DNA. There is some evidence that this unusual component is a mechanism to evade bacterial defense mechanisms such as restriction endonucleases and CRISPR/Cas systems which evolved to recognize and cleave sequences within invading phages, thereby inactivating them. Other phages have long been known to use unusual nucleotides. In 1963, Takahashi and Marmur identified a Bacillus phage that has dU substituting dT in its genome, and in 1977, Kirnos et al. identified a cyanophage containing 2-aminoadenine (Z) instead of adenine (A).
Systems biology
The field of systems biology investigates the complex networks of interactions within an organism, usually using computational tools and modeling. For example, a phage genome that enters into a bacterial host cell may express hundreds of phage proteins which will affect the expression of numerous host genes or the host's metabolism. All of these complex interactions can be described and simulated in computer models.
For instance, infection of Pseudomonas aeruginosa by the temperate phage PaP3 changed the expression of 38% (2160/5633) of its host's genes. Many of these effects are probably indirect, hence the challenge becomes to identify the direct interactions among bacteria and phage.
Several attempts have been made to map protein–protein interactions among phage and their host. For instance, bacteriophage lambda was found to interact with its host, E. coli, by dozens of interactions. Again, the significance of many of these interactions remains unclear, but these studies suggest that there most likely are several key interactions and many indirect interactions whose role remains uncharacterized.
Host resistance
Bacteriophages are a major threat to bacteria and prokaryotes have evolved numerous mechanisms to block infection or to block the replication of bacteriophages within host cells. The CRISPR system is one such mechanism as are retrons and the anti-toxin system encoded by them. The Thoeris defense system is known to deploy a unique strategy for bacterial antiphage resistance via NAD+ degradation.
Bacteriophage–host symbiosis
Temperate phages are bacteriophages that integrate their genetic material into the host as extrachromosomal episomes or as a prophage during a lysogenic cycle. Some temperate phages can confer fitness advantages to their host in numerous ways, including giving antibiotic resistance through the transfer or introduction of antibiotic resistance genes (ARGs), protecting hosts from phagocytosis, protecting hosts from secondary infection through superinfection exclusion, enhancing host pathogenicity, or enhancing bacterial metabolism or growth. Bacteriophage–host symbiosis may benefit bacteria by providing selective advantages while passively replicating the phage genome.
In the environment
Metagenomics has allowed the in-water detection of bacteriophages that was not possible previously.
Also, bacteriophages have been used in hydrological tracing and modelling in river systems, especially where surface water and groundwater interactions occur. The use of phages is preferred to the more conventional dye marker because they are significantly less absorbed when passing through ground waters and they are readily detected at very low concentrations. Non-polluted water may contain approximately 2×108 bacteriophages per ml.
Bacteriophages are thought to contribute extensively to horizontal gene transfer in natural environments, principally via transduction, but also via transformation.Metagenomics-based studies also have revealed that viromes from a variety of environments harbor antibiotic-resistance genes, including those that could confer multidrug resistance.
In humans
Although phages do not infect humans, there are countless phage particles in the human body, given our extensive microbiome. Our phage population has been called the human phageome, including the "healthy gut phageome" (HGP) and the "diseased human phageome" (DHP). The active phageome of a healthy human (i.e., actively replicating as opposed to nonreplicating, integrated prophage) has been estimated to comprise dozens to thousands of different viruses. There is evidence that bacteriophages and bacteria interact in the human gut microbiome both antagonistically and beneficially.
Preliminary studies have indicated that common bacteriophages are found in 62% of healthy individuals on average, while their prevalence was reduced by 42% and 54% on average in patients with ulcerative colitis (UC) and Crohn's disease (CD). Abundance of phages may also decline in the elderly.
The most common phages in the human intestine, found worldwide, are crAssphages. CrAssphages are transmitted from mother to child soon after birth, and there is some evidence suggesting that they may be transmitted locally. Each person develops their own unique crAssphage clusters. CrAss-like phages also may be present in primates besides humans.
Commonly studied bacteriophage
Among the countless phage, only a few have been studied in detail, including some historically important phage that were discovered in the early days of microbial genetics. These, especially the T-phage, helped to discover important principles of gene structure and function.
See also
- Bacterivore
- CrAssphage
- CRISPR
- DNA viruses
- Macrophage
- Phage ecology
- Phage monographs (a comprehensive listing of phage and phage-associated monographs, 1921–present)
- Phagemid
- Polyphage
- RNA viruses
- Transduction
- Viriome
- Virophage, viruses that infect other viruses
Bibliography
- Hauser AR, Mecsas J, Moir DT (July 2016). "Beyond Antibiotics: New Therapeutic Approaches for Bacterial Infections". Clinical Infectious Diseases. 63 (1): 89–95. doi:10.1093/cid/ciw200. PMC 4901866. PMID 27025826.
- Strathdee S, Patterson T (2019). The Perfect Predator. Hachette Books. ISBN 978-0316418089.
External links
- Häusler T (2006). Viruses vs. superbugs : a solution to the antibiotics crisis?. London: Macmillan. ISBN 978-1-4039-8764-8.
- Abedon ST. "The Bacteriophage Ecology Group". The Ohio State University. Archived from the original on 3 June 2013.
- Tourterel C, Blouin Y. "Bacteriophages illustrations and genomics". Orsay phage web site. Archived from the original on 29 October 2013. Retrieved 24 October 2013.
- "QuipStories: Bacteriophages get a foothold on their prey" (PDF). PDBe.
- Flatow I (April 2008). "Using 'Phage' Viruses to Help Fight Infection". Science Friday podcast. NPR. Archived from the original on 17 April 2008.
- "Animation of a scientifically correct T4 bacteriophage targeting E. coli bacteria". YouTube.
- "T4 Bacteriophage targeting E. coli bacteria". Animation by Hybrid Animation Medical. 21 December 2009.
- Bacteriophages: What are they. Presentation by Professor Graham Hatfull, University of Pittsburgh on YouTube
| Components | ||
|---|---|---|
| Viral life cycle | ||
| Genetics | ||
| By host | ||
| Other | ||